Replicate
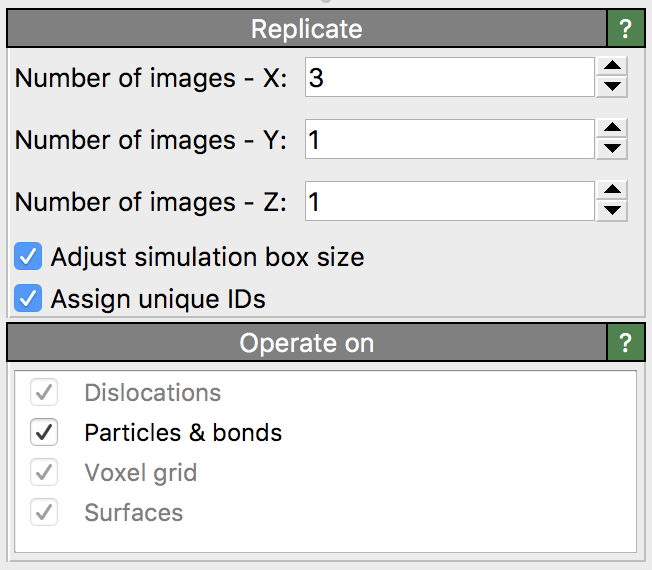
This modifier copies all particles, bonds, and other data elements multiple times to visualize periodic images of a system.
The Operate on list in the lower panel lets you select the kinds of objects that should be replicated by the modifier. By default, the modifier extends the simulation cell appropriately to encompass all generated images of the system. If not desired, you can turn off the option Adjust simulation box size to keep the original simulation cell geometry. You should be aware, however, that this produces an inconsistent state, where the periodicity length no longer fits the explicitly replicated contents of the simulation cell.
Parameters
- Number of images - X/Y/Z
These values control how many times the system is copied along each simulation cell vector.
- Adjust simulation box size
Extends the simulation cell to match the size of the replicated system.
- Assign unique IDs
This option lets the modifier assign new unique IDs to the copied particles or bonds. Otherwise, the duplicated elements will have the same identifiers as the originals, which may cause problems with other modifiers (e.g. the Manual selection modifier), which rely on the uniqueness of identifiers.
For replicated particles, this option also adjusts their
Molecule Identifierproperty if it exists.
Added in version 3.10.1: If the Periodic Image particle property is present and Adjust simulation box size is not turned off, the modifier will implicitly
un-wrap the particle coordinates before replicating the system and then re-wrap the
coordinates of the particles again. This ensures that molecule identifiers assigned to
particles remain correct if the molecules wrap around at periodic cell boundaries.
For further information, see also the notes on image flags in the docs of the LAMMPS replicate command.
See also
ovito.modifiers.ReplicateModifier (Python API)